Evolution of shape-defined macromolecules into functional systems
SHAPE
Functionalities of enzymes are encoded in amino acid sequences and directed by their SHAPEs with complementary binding pockets for specific substrates. Natural enzymes are remarkable catalysts, however, they are typically optimized by evolution to operate under the constraints of the physiological environment of a living system, which strongly limits the scope of their applications in organic synthesis. Here, I propose to develop abiotic enzymes to selectively catalyze chemical transformations in non-physiological environments. The main objective of the project is to use monomer sequence control to fine-tune the SHAPE of abiotic macromolecules to obtain the desired catalytic functionality. Our approach proposes an unexplored method for obtaining abiotic, sequence-defined polymers whose functions can rival those of natural macromolecules. The study will reveal valuable information on sequence-dependent properties of polymers, to open a field of abiotic enzymes for organic transformations.

Project ERC Starting Grant No 101116700
Sequence programmable polymer materials for the selective sensing of bioactive water pollutants PolySens
PolySens
The problem of water pollution with bioactive compounds has been growing in the last few years due to the increasing consumption of drugs in developed societies and the pollution of the environment with plastics. Drugs we take are not fully metabolized, are excreted by our body and end up in the natural environment. In particular, very dangerous are endocrine-disrupting chemicals (EDCs), as pointed out by the World Health Organization. EDCs can affect the endocrine system and subsequently impair the development and fertility of animals and humans. The aim of the project is to develop a highly sensitive and selective method of detecting bioactive water pollutants based on fluorescent polymers with a defined structure. The monomer sequence of the polymer will be designed to specifically interact with the selected analytes. Programming polymer properties through sequences is a breakthrough approach in the field of polymer materials and allows for very precise modulation of their properties. Polymers defined in their structure can perform functions like natural macromolecules, e.g. receptors while maintaining appropriate material properties, i.e. stability. Providing tools for efficient control of the level of bioactive pollutants in water will allow for the widespread introduction of water quality monitoring programs. Wastewater treatment plants will be the direct recipients of the project results and indirectly the whole community.

Asymmetric catalysis using enzyme-like ligands based on sequence-programmable oligourethanes
PolyCat
Nature has encoded the secret of life in a sequence of biopolymers such as nucleic acid and proteins. For example, enzymes are capable of catalyzing several biochemical reactions with absolute selectivity. This is attributed to the 3-dimensional geometry of enzymes which determines their functions and activity. Recapitulating similar functions in synthetic macromolecules is a challenge in modern polymer chemistry. The non-natural sequence-defined polymers have great potential to exhibit self-assembly and programmed folding and received widespread attention in material and life sciences. However, there is limited prior information on inducing enzyme-like catalytic properties of synthetic sequence-defined polymers for abiotic chemical transformations. Therefore, we aim to investigate sequence-defined oligourethanes as ligands in asymmetric catalysis. The project will add fundamental knowledge on the synthesis and conformational characteristics of stereo-controlled sequence-defined polyurethanes. This understanding of sequence-structure correlation will further be applied to develop potential ligands for catalytic hydrogenation of alkenes with high selectivity.
Project is carried out in collaboration with Dr Pawel Dydio, Complex Systems in Synthesis & Catalysis Laboratory at ISIS, Université de Strasbourg, Centre national de la recherche scientifique, France.

Sequence-regulated polymer self-assembly towards materials mimicking living systems
MimicLS
In living matter, self-assembly benefits from the evolutionary processes that tune interactions to optimize the properties, morphology and functionality of the resulting biomaterials. Nature exploits the primary sequence of natural macromolecules (e.g., proteins) that fold into particular motifs that self-organize into complex structures representing various properties. In contrast, man-made polymer materials are far away from advanced functionalities as represented by living systems.
Nowadays, the progress in polymer synthesis enables full control of monomer sequences with biological precision. However, to enable their practical use a sustainable and highly efficient approach has to be developed. It is expected that sequence-defined macromolecules can be designed to fold into particular 3D structures by a selection of the proper monomer alphabet, as it is observed for natural macromolecules. Yet, very little is known about single chain folding of non-natural macromolecules with defined primary structure and their assembly into complex supramolecular structures has not been investigated, so far.
The project will be carried out in cooperation between:
(i) Roza Szweda`s Group, Poland, an expert in precision polymer chemistry,
(ii) Takuji Adachi`s Group from the Faculty of Sciences at the University of Geneva, Switzerland, physical chemist and specialist in optical spectroscopy on self-assembled materials, developing in-situ spectroscopy tools,
We jointly took a challenge to investigate the fundamental self-assembly process of abiotic, sequence-defined polymers. The project aims to obtain knowledge on sequence-regulated, hierarchical polymer self-assembly, which is required for creating synthetic materials with structural sophistication and complex function as represented by living matter.


Programmable materials based on polymers of defined monomer sequence
PolyMat
Inspired by natural macromolecules, sequence‐defined synthetic polymers are promising materials building blocks that integrate the sequence‐derived activity of natural biopolymers with the robustness and versatility of synthetic systems. Sequence-defined macromolecules are components of natural matter, that create living objects able to move, work and even think. In spite of many efforts, man-made materials are still far away from functions that are displayed by natural matter. To reach for the diverse structures and complex properties achieved by native biological polymers sequence programmability of the synthetic polymer is required.
Recently, we have developed a new synthesis method for sequence-defined polymers, based on system chemistry approaches. The method has many advantages over existing methods such as scalability, low cost, reduced amount of solvents, and easy protocol. This innovative solution has opened the door to large-scale synthesis of polymers with a specific sequence and enabled their use as building blocks in materials. In this project based on our invention, we are going to develop new polymer materials built from sequence-defined polymers with finely tunable properties. The properties of the materials are programmed by monomer sequence.
Polymers as a next generation data storage media PolyDigit
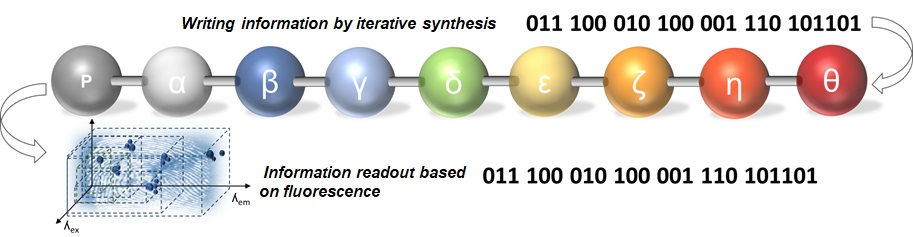
Nature uses DNA for data storage – natural polymers made of nucleotides that contain all the information necessary for the proliferation of living organisms. Defined Sequence Polymers (SDPs) offer a stable, resource-efficient, energy-efficient, and sustainable data storage solution. The properties of synthetic polymers can be precisely modulated and adapted to the requirements. The characteristics of the polymer can be fine-tuned by selecting appropriate building blocks from a wide library of synthetic monomers. Modern, advanced nanosynthesis methods allow materials to be easily produced with the precision of monolayers. Macromolecules can be sequentially arranged in a small volume and used as high-capacity data storage materials. Currently, the challenge is to read the information stored in digital polymers easily and quickly. The lack of convenient methods for reading information using non-destructive techniques prevents the development of new data storage materials based on synthetic polymers. The project aims to study fluorescent polymer materials for their usefulness in storing data and reading information encoded in the monomer sequence based on the fluorescence pattern.
Funding:

Sequence-defined macromolecules of controlled folding ConFold
The natural, uniform macromolecules such as proteins and DNA of sequence-defined structures have been inspiring polymer chemists for years. The roles and functions that they can attain are determined by their three-dimensional arrangement that depends on the monomer sequence. To reach for the diverse structures and complex properties, represented by native biological polymers, sequence programmability of the synthetic polymer and control of their 3D structure are required.

ConFold will add a fundamental knowledge of synthesis and structural properties of synthetic sequence-defined polymers to fill a part of the large information gap concerning properties and displayed functions between synthetic polymers and native biological materials. This understanding of sequence-structure relationships will enable us to gain better control over polymers properties and will advance their application scope.
